
For those having the same “contig name” will be grouped into the same contig. Then, sangeranalyseR uses the str_split function to split and vectorize their filenames into “contig name” and “direction-suffix” two parts. So how sangeranalyseR works is that it first matches the forward and reverse reads by matching REGEX_SuffixForward and REGEX_SuffixReverse. Here is an example of how it works in sangeranalseR: Please refer to What is a regular expression? for more details. If you don’t know what regular expression is, don’t panic - it’s just a way of recognising text.
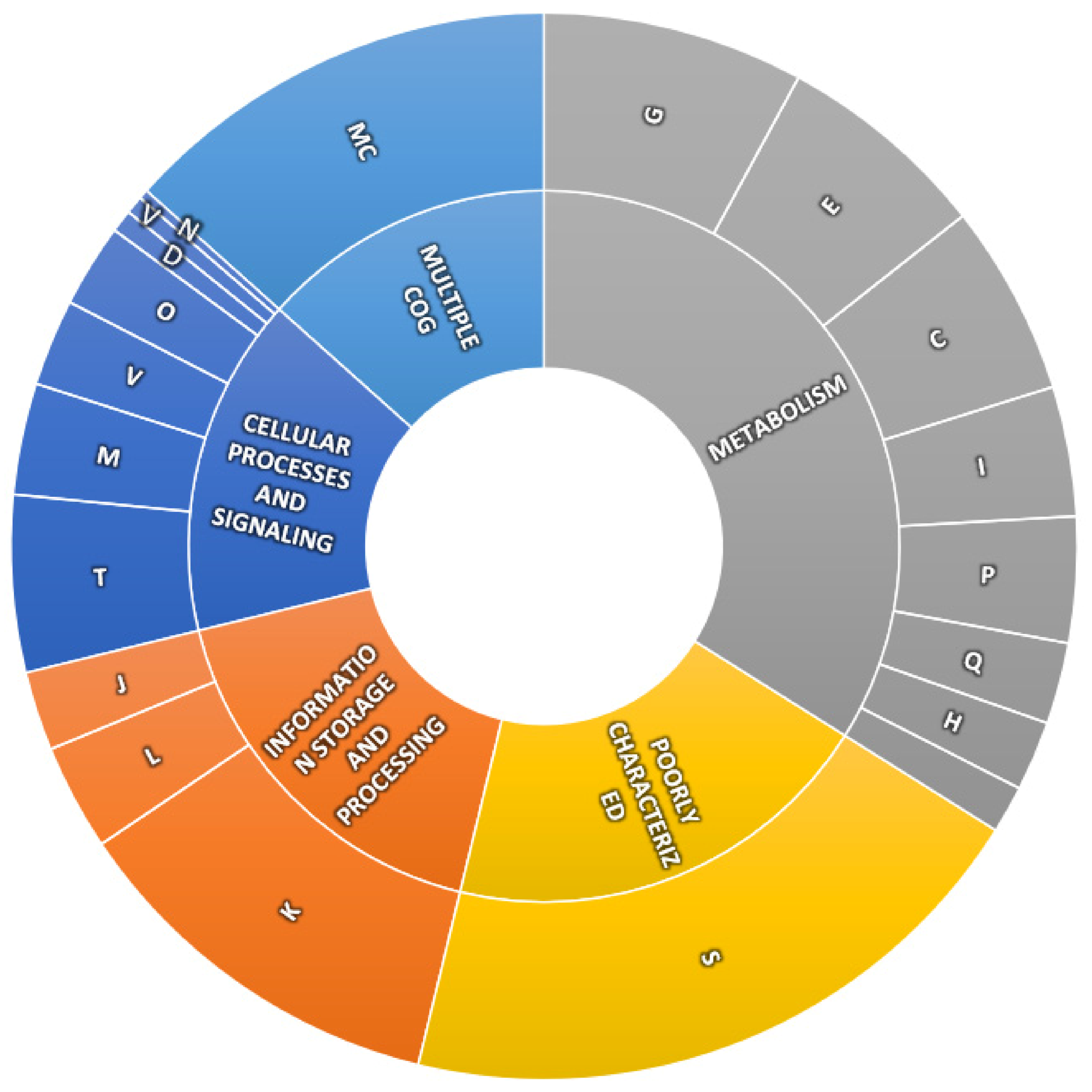

(2) Creating SangerAlignment instance from AB1.(1) Preparing SangerAlignment AB1 inputs.Generating SangerAlignment report (AB1).Writing SangerAlignment FASTA files (AB1).Updating SangerAlignment quality trimming parameters.(2) “CSV file matching” SangerAlignment creation ( AB1).(1) “regular expression matching” SangerAlignment creation ( AB1).Creating SangerAlignment instance from AB1.(2) “CSV file matching” SangerAlignment inputs ( AB1).(1) “regular expression matching” SangerAlignment inputs ( AB1).Advanced User Guide - SangerAlignment ( AB1).Advanced User Guide - SangerContig ( AB1).Advanced User Guide - SangerRead ( AB1).Their ability to perform aerobic respiration is evidenced by the presence of both cytochrome d complex and all subunits of proton-translocating NADH-dehydrogenase genes (nuoA-N). This process was iterated as needed to close gaps that could not be closed using the De Brujin graph method.īoth genomes were automatically annotated by the PGAP pathway at NCBI, and the resulting open reading frames were compared with RAST ( 3) and KEGG annotations ( 4). Gaps were closed using BioEdit ( 2) based on alignments of reads: reads corresponding to the ends of each contig were located, sorted, and aligned using JMP version 10 (SAS Institute Inc., USA), and then visualized using BioEdit.
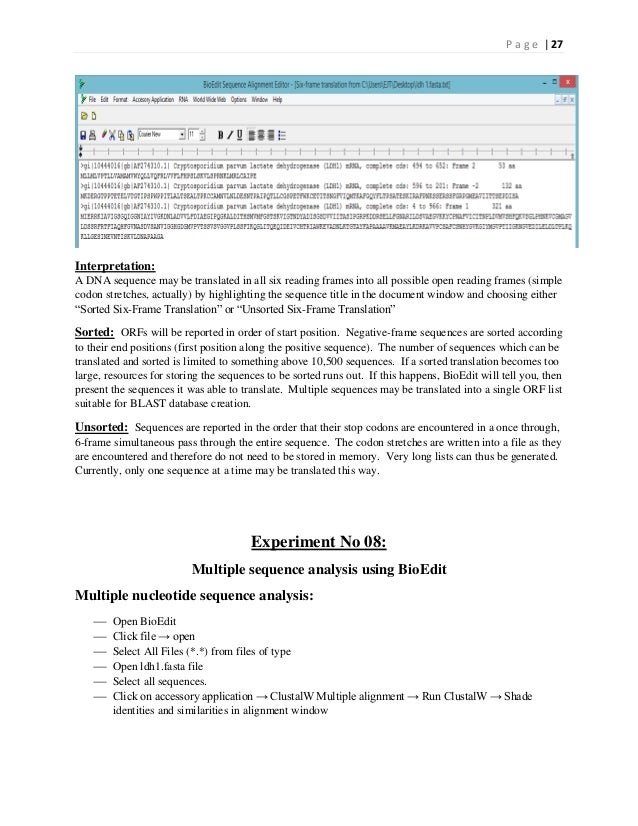
#Change nuclotide contig in bioedit software
Reads were trimmed based on quality (limit = 0.05), and assembled using the De Bruijn graph method of de novo assembly provided by CLC genomics workbench version 7.04, with default parameters.ĪflIII (both strains) and HinDII (X1698 only) optical maps were analyzed using Argus MapSolver software (OpGen, USA) to order and orient contigs with respect to each other. Sequence reads were 100-bp × 100-bp paired-end reads on a HiSeq2500 operating in rapid mode 12,496,108 (X1036 T) and 12,804,044 (X1698) sequence reads were generated. Genome libraries were prepared using the NEB Ultra DNA library prep kit (New England Biolabs, USA), according to the manufacturer’s instructions, on a PerkinElmer Sciclone NGS robot.
Cells were harvested and gDNA was prepared using the Epicentre Metagenomic DNA isolation kit for water (Illumina, USA). Genomic DNA was extracted from cells grown on CDC anaerobic blood agar (BD) in a GasPak for 2 to 3 weeks. (GasPak and CampyPak are products of Becton, Dickinson and Company, USA.) Limited growth is achieved under the conditions created by a CampyPak (6 to 16% O 2) and little to no growth in ambient air. The strains grow best in an atmosphere generated by a GasPak in an anaerobe jar (≤1% O 2), including the stringent environment of an anaerobe chamber. Two strains of a novel fastidious, partially acid-fast, Gram-positive Corynebacterineae bacterium (X1036 T and X1698) were obtained from human abscesses, as previously described ( 1). The Special Bacteriology Reference Laboratory receives unusual and difficult-to-identify bacterial strains derived from clinical specimens from throughout the United States.
